Group Equivariant Convolutional Networks in Medical Image Analysis
Introduction
AI has been one of the most heated topic in this planet recently and greatly changed people's life. Past years has witnessed the widely applications of convolutional neural networks (CNNs) in medical image analysis, including several tasks such as classification, segmentation, which greatly advanced diagnosis and treatment in clinical practice.
CNNs' success rely on reasonable architectures of networks, and strong computational power, as well as large scale datasets. But in some scenarios like healthcare system, large scale data is hard to obtain. Thus extensive data augmentation should be implemented to achieve better performance.
Different from natural images, some medical images exhibit not only translational symmetry but rotation and reflection symmetry. However, CNNs failed to make good use of these symmetries.
Group equivariant convolutional networks (G-CNNs) [1] was proposed in 2016 as a generalization of CNNs, using G-convolutions to enjoy a substantially higher degree of weight sharing, which can be implemented with negligible computational overhead for discrete groups generated by translations, reflections and rotations. This architecture has been used in medical image analysis and proved that it can improve the performance in multi-tasks
This paper will give a brief review for the applications of G-CNNs in medical image analysis and show some details about how it improves the models.
Prelimiaries
Symmetries
Cohen et al. [1:1] explained the concept of symmetry as transformation that maintains the object invariant or equivalent. Cohen's PhD thesis [2] further explained that the set of all symmetries can be considered as a label function where the equivalence classes induced by the group are the same as the classes.

Figure 1: Example of a label function
Figure 1 [2:1] is an example of a label function
As
And [1:2] also takes sampling grid of images as an example, which is easier to understand: for a sampling grid of images
We call a set of transformations with properties of symmetry as symmetry group.
Representations
After the introduction symmetry, let us go to "representations". When the symmetry transformations act on vectors, we have a representation of the group. In [2:2], group representation is defined as the appropriate notion of group.
To be easier, a group action
Feature maps of CNNs as spaces of functions can also be regarded as one kind of representation.
Equivariance
Equivariance is a kind of important symmetry.
Define
For
where
[2:3] also pointed out that equivariance is transitive: if
In machine learning, generally, equivariance is of more importance than invariance, as we can judge spatial relationships based on equivariant features rather than invariant features.
The group
[1:3] proposed the concept of two types of groups,
where
The group
where
Group Equivariant Convolutional Networks
Structure of Symmetric Feature Maps
The map from images to stacks of feature maps with
A transformation
Feature maps in a
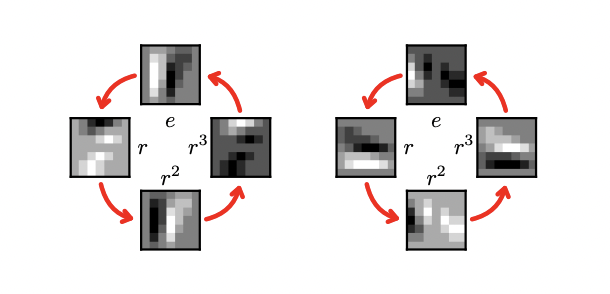
Figure 2: A p4 feature map and its rotation by r

Figure 3: A p4m feature map and its rotation by r
The Structure of Group Equivariant Convolutional Networks
The first layer of Group Equivariant Convolutional Networks is used to lift the images (a sampling grid of images
And all layers after the first can be defined as:
The difference is that the function is on the plane
Both these two functions are proved to be equivariant in [1:5].
3D Group Convolutions
[3] proposed a 3D Group Convolution to solve 3D images' analysis like CT scans. The 3D group convolution is implemented as
where
The
Applications in Medical Image Analysis
Classification
General process of classification consists of three steps by using G-CNNs:
- Convert an image to feature maps
- Convert feature maps to feature maps
- Convert feature maps to a label.
In the final layer, a group-pooling layer is used to ensure that the output is invariant as a function on the plan for classification tasks.[4]
Example1: [4:1] proposed equivariant DenseNet architecture for the
- Define the G-convolution on image
-convolution:
where
- Define the
-convolution:
- Define a projection layer:
The final layer of the model is a
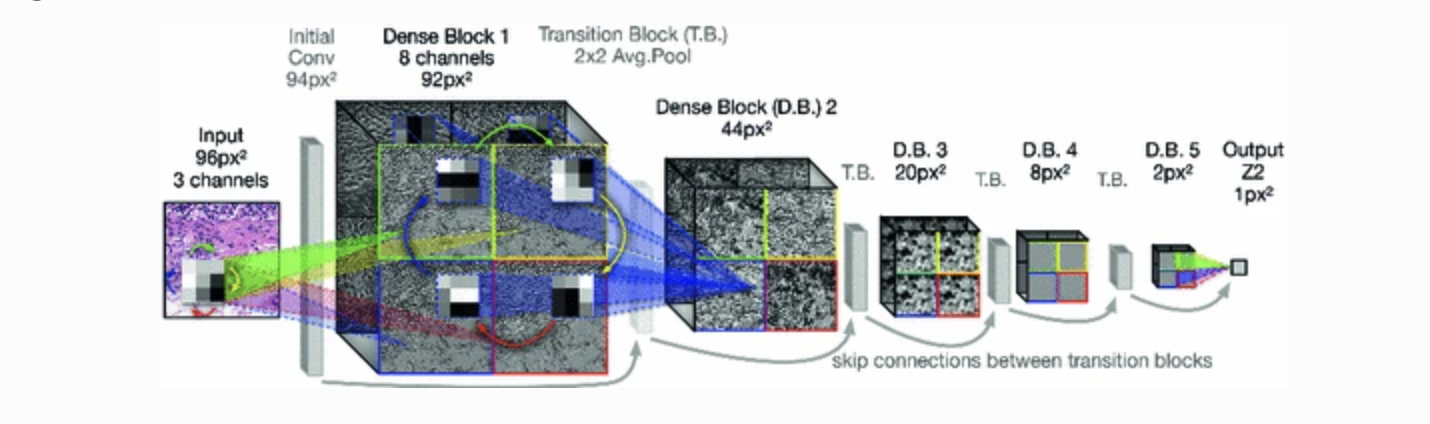
Figure 4: Equivariant DenseNet architecture, the figure is from [^5]
Example2: [5] proposed a kind of dynamic group convolution to improve the performance of breast tumor classification.The dynamic group convolutional is parameterized as a dynamic combination rather than a fixed one of multiple kernels. And the operation was generalized to an equivariant one.
For
where
The paper proved that the dynamic group convolution operator's equivariance can be maintained by giving an equivariant constraint on
The results and comparisons on PCAM Dataset is shown as below, which proved that dynamic group convolution can led to improvement in breast tumor classification:
| Method | Group | AUC | |
| DenseNet | |||
| DenseNet+ | |||
| P4-DenseNet | |||
| P4M-DenseNet | |||
| Dy-DenseNet | |||
| Dy-DenseNet+ | |||
| Dy-P4-DenseNet | |||
| Dy-P4M-DenseNet |
Example3: [3:1] applied 3D GCNNs on Nodule classification and evaluated performance by using Free-Response Operating Characteristic (FROC)[].
The results of and comparison is shown as below, from which we can see the 3D G-CNNs can improve classification performance:
| 30 | |||||
| 300 | |||||
| 3,000 | |||||
| 30,000 |
Segmentation
General process of segmentation consists of three steps by using G-CNNs:
- Convert an image to feature maps,
- Convert feature maps to feature maps
- Convert feature maps to a segmentation.
In the final layer, a group-pooling layer is used to ensure equivariant as a function on the plane (for segmentation tasks, where the output is supposed to transform together with the input).[4:2]
Example1: [6] proposed a Rota-Net to segment simultaneous gland and lumen segmentation in colon histology images, incorporating the inherent rotational symmetry within histology images into an encoder-decoder based network by utilizing G-CNNs, specifically using the symmetry group of rotations by multiples of 90∘.
The main process is as below:
- Define the
-convolution on image
where
- Define the
-convolution on feature maps as:
- Define the projection layer:
The architecture of Rota-Net is as the figure [6:1]:
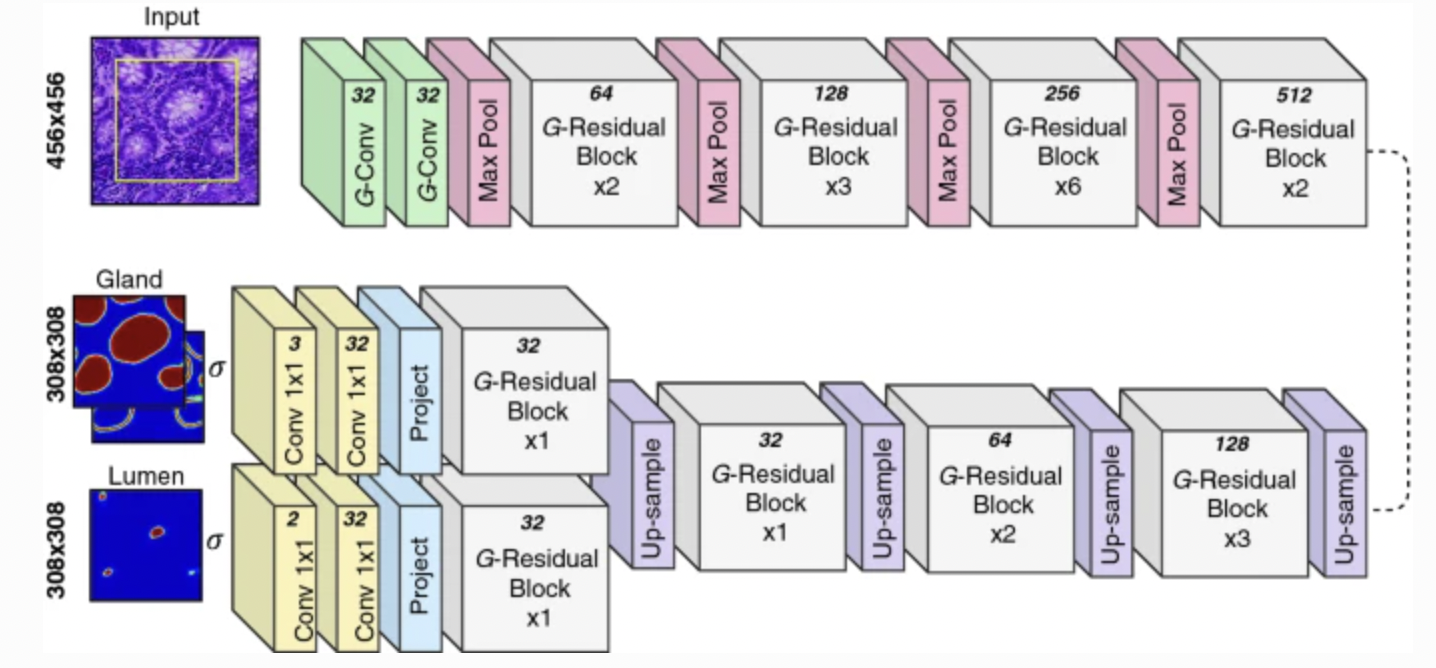
Figure 5: The architecture of Rota-Net: the yellow box within the input means the part of the image considered at the output. The number at the top of each operation means the number of feature maps produced.
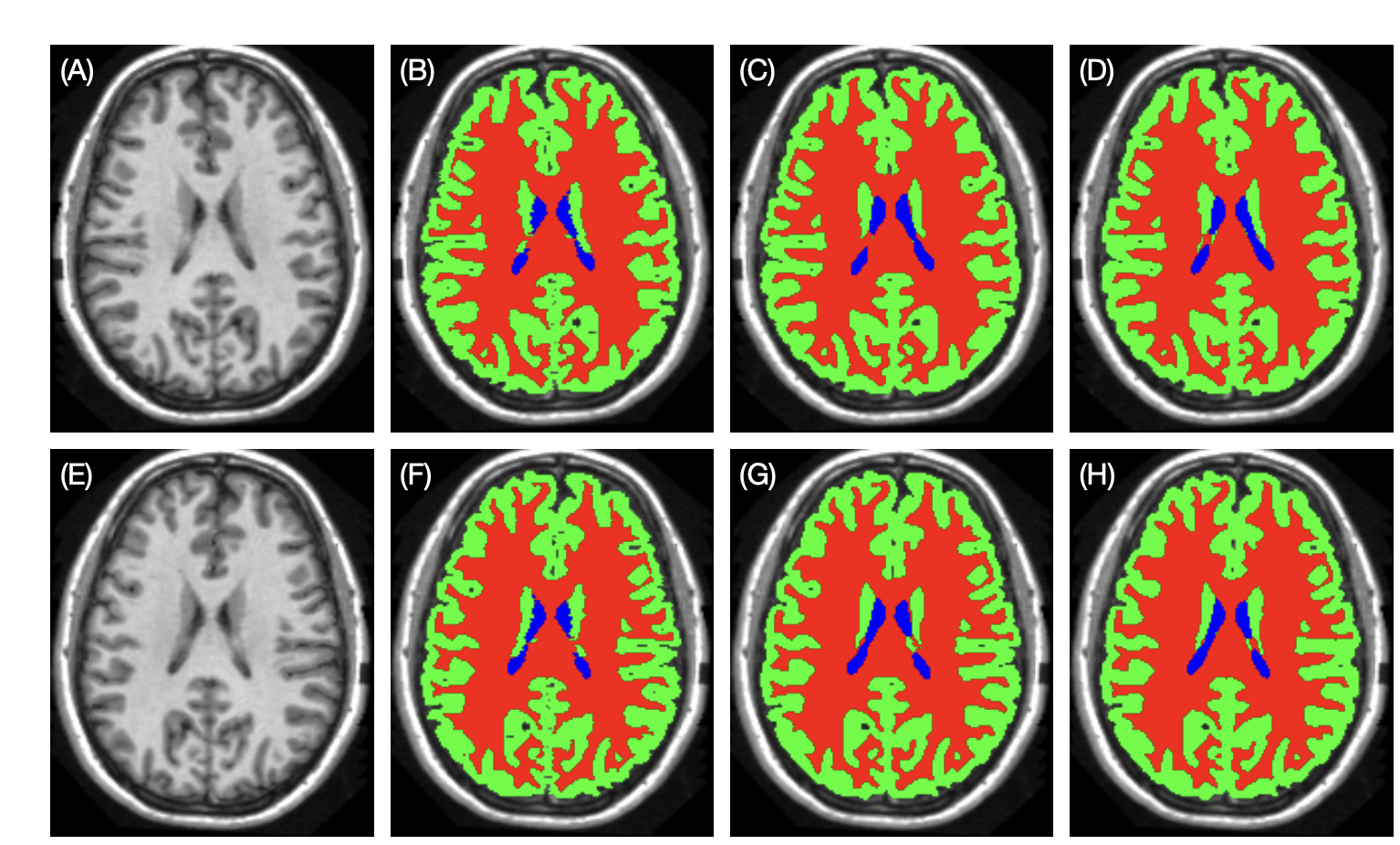
Example2: [7] propose a 3D left-right-reflection equivariant network to segment the anatomical structures of the brain, exploiting the left and right symmetry property of the brain. The segmentation model was 3D U-Net[8], which is a classical segmentation model and it incorporated RE convolutions.
For a multi-channel function
The difference between in G-CNNs between example 1 and example 2 is that the feature map is 3D, and the transformation is left-right-reflection. The comparison of result is shown as below [7:1]:
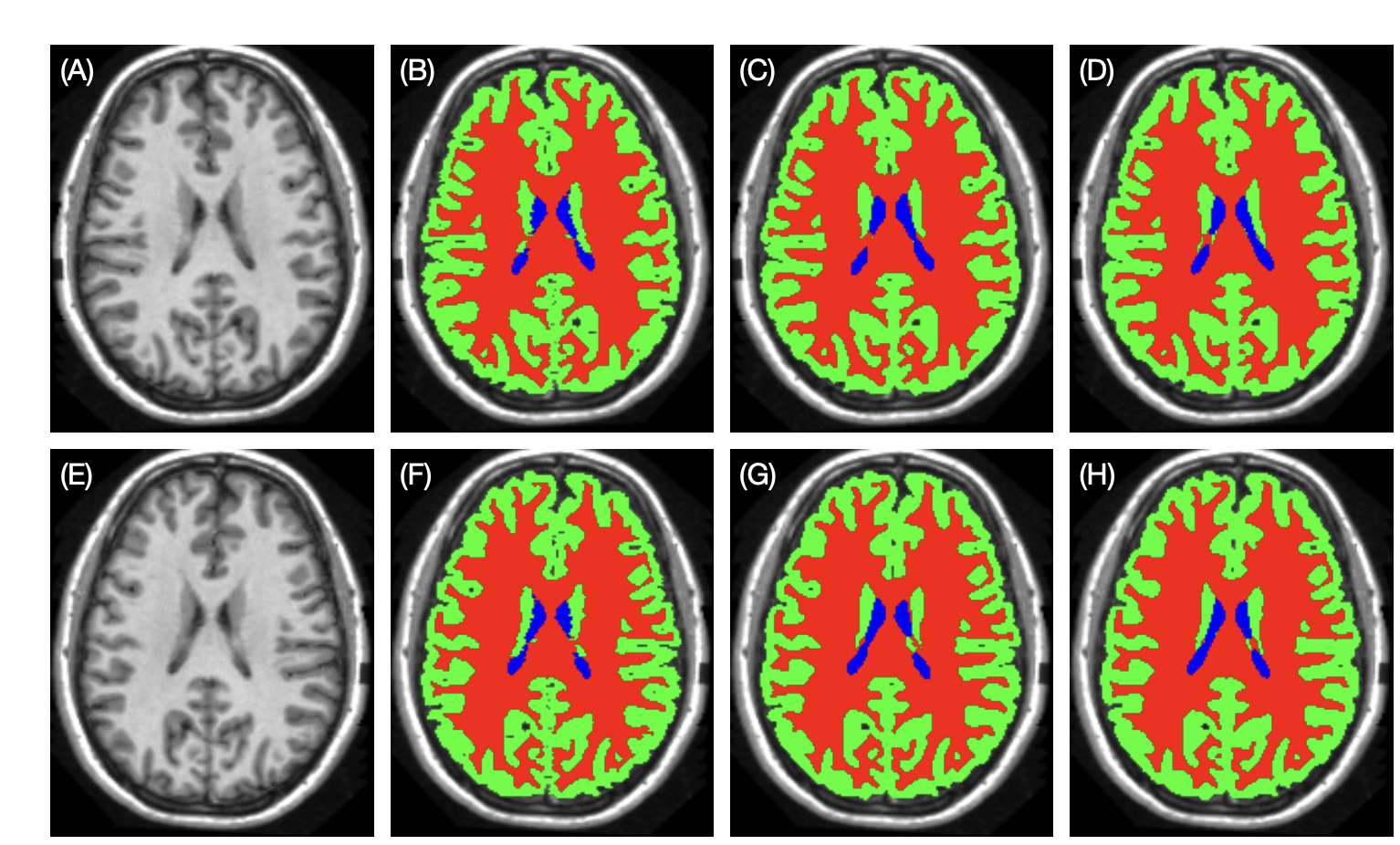
Figure 6: Example brain tissue segmentation. (A), (E): original and reflected testing images. (B), (F): manual delineations. (C), (G): results of the conventional U-Net trained with reflection augmentation. (D), (H): the results of the RE U-Net.
Example3: [9] proposed a new model of two-pathway-group CNN architecture for brain tumor segmentation, making use of local features and global contextual features at the same time. Equivariance has been maintained to reduce instabilities and overfitting parameter sharing, which could improve the segmentation performance. The following figure is from [9:1]
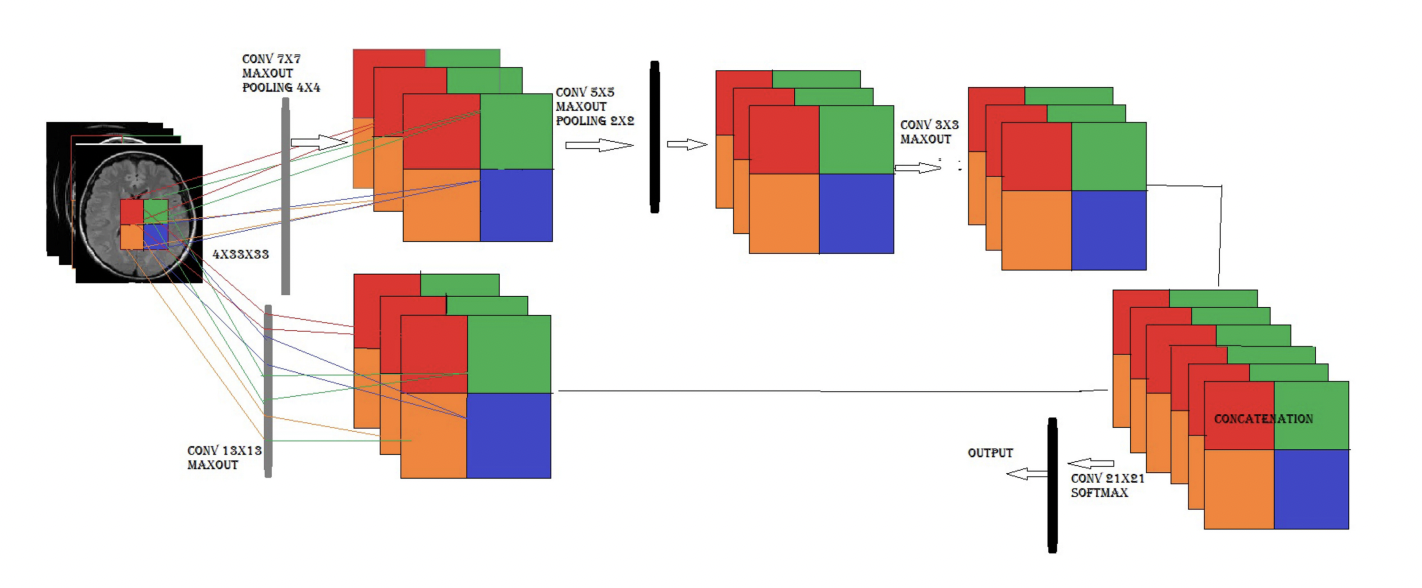
Figure 7: Two-Pathway-Group CNN architecture (2PG-CNN) showing that the input patch is processed by two different group CNNs. The four blocks in a feature map of both CNNs show a p4 group features map that inherits group CNN properties. The upper CNN represents a local feature map and the lower CNN shows a global feature map.
Conclusions and Future Opportunities
This paper reviews the basic theory of G-CNNs, and gives some typical examples of its application on medical images. Due to the characteristics of lesions, tissues or structures, medical images are very suitable for using G-CNNs. It can be seen from many examples that compared with the standard CNN, G-CNNs can effectively improve the performance of the model as a more generalized one.
In future studies, we can design the combination of G-CNNs and classification or segmentation models according to the specific characteristics of lesions, tissues, structures due to the more complex symmetries in 3D images. In addition, we can also analyze the latest state-of-the-arts models and explore the possibility of combining with G-CNNs.
Besides, the examples in this review focus on classification and segmentation, but there are also tasks of generation and reconstruction in medical image analysis. Equavariant neural networks can also combine with methods in these tasks. Additionally, frame equavariance may be used in clinical videos analysis.
And the equavariance not only can be explored in CNNs, but also in Transformers. As the great success of transformers has achieved these days, there would also are opportunities for the research of Transformers and their exploiting with equavariance, as well as potential applications in medical image analysis.
Reference
- Cohen, T. S. (2021). Equivariant convolutional networks. (https://dare.uva.nl/search?identifier=0f7014ae-ee94-430e-a5d8-37d03d8d10e6) ↩︎ ↩︎ ↩︎ ↩︎
- Van Ginneken, B., Armato III, S. G., de Hoop, B., van Amelsvoort-van de Vorst, S., Duindam, T., Niemeijer, M., ... & Prokop, M. (2010). Comparing and combining algorithms for computer-aided detection of pulmonary nodules in computed tomography scans: the ANODE09 study. Medical image analysis, 14(6), 707-722.(https://www.sciencedirect.com/science/article/pii/S1361841510000587) ↩︎ ↩︎
- Veeling, B. S., Linmans, J., Winkens, J., Cohen, T., & Welling, M. (2018, September). Rotation equivariant CNNs for digital pathology. In International Conference on Medical image computing and computer-assisted intervention (pp. 210-218). Springer, Cham (https://link.springer.com/chapter/10.1007/978-3-030-00934-2_24) ↩︎ ↩︎ ↩︎
- Li, Y., Cao, G., & Cao, W. (2020, December). A dynamic group equivariant convolutional networks for medical image analysis. In 2020 IEEE International Conference on Bioinformatics and Biomedicine (BIBM) (pp. 1056-1062). IEEE.(https://ieeexplore.ieee.org/abstract/document/9313601) (https://www.sciencedirect.com/science/article/pii/S1361841510000587) ↩︎
- Graham, Simon, David Epstein, and Nasir Rajpoot. "Rota-net: Rotation equivariant network for simultaneous gland and lumen segmentation in colon histology images." European Congress on Digital Pathology. Springer, Cham, 2019. (https://link.springer.com/chapter/10.1007/978-3-030-23937-4_13) ↩︎ ↩︎
- Han, Shuo, Jerry L. Prince, and Aaron Carass. Reflection-equivariant convolutional neural networks improve segmentation over reflection augmentation.Medical Imaging 2020: Image Processing. Vol. 11313. International Society for Optics and Photonics, 2020. (https://www.spiedigitallibrary.org/conference-proceedings-of-spie/11313/1131337/Reflection-equivariant-convolutional-neural-networks-improve-segmentation-over-reflection-augmentation/10.1117/12.2549399.full) ↩︎ ↩︎
- Çiçek, Ö., Abdulkadir, A., Lienkamp, S. S., Brox, T., & Ronneberger, O. (2016, October). 3D U-Net: learning dense volumetric segmentation from sparse annotation. In International conference on medical image computing and computer-assisted intervention (pp. 424-432). Springer, Cham. (https://link.springer.com/chapter/10.1007/978-3-319-46723-8_49) ↩︎
- Razzak, M. I., Imran, M., & Xu, G. (2018). Efficient brain tumor segmentation with multiscale two-pathway-group conventional neural networks. IEEE journal of biomedical and health informatics, 23(5), 1911-1919.(https://ieeexplore.ieee.org/abstract/document/8481481) ↩︎ ↩︎